EasyModeller 4.0 introduces a fresh new GUI for Homology Modelling using MODELLER in the backend and available for both Windows and Linux platform. This version has several new features integrating all the goodies of EasyModeller 3.0 which was only available for LINUX.
The highlighted features of this version are :
1. Tab based logical Modelling steps with extensive error handling
2. Allows to load unlimited number of templates
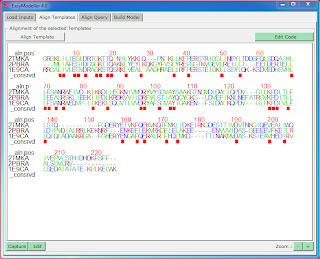
3. A colorful alignment viewer and also with an inbuilt alignment editor
4. On the fly MODELLER code editing ( generated MODELLER scripts can be edited as per user's need and run from within the tool). This feature would be most useful for advanced MODELLER users.
5. Inbuilt DOPE profile Viewer, Ramachandran Plot viewer, Loop Modelling, Basic model optimization and dynamics for a selected Model.
DOWNLOAD Link : Click here
If you use EasyModeller in your work please cite http://www.ncbi.nlm.nih.gov/pubmed/20712861
Before using EasyModeller please make sure that :
1. You have downloaded the correct & latest version of MODELLER and installed the same in your PC ( Please follow the installation guides )
2. You have entered the correct license key for MODELLER ( obtained free after registering)
3. You have downloaded and installed a compatible Python version ( preferably 2.6 or 2.7 but not >=3)
It would be best that if you first test whether MODELLER is working correctly in your system. To do this follow the steps below :
1. Go to command prompt (or terminal) and type :python
This should open the python IDLE and you should see a message like this :
Python 2.7.1+ (r271:86832, Apr 11 2011, 18:05:24)
[GCC 4.5.2] on linux2
Type "help", "copyright", "credits" or "license" for more information.
>>>
N.B : If you have installed Python and still do not see this then make sure that you have set your path for Python correctly (although this should be done automatically)
2. Now inside the Python IDLE type : import modeller
Python 2.7.1+ (r271:86832, Apr 11 2011, 18:05:24)
[GCC 4.5.2] on linux2
Type "help", "copyright", "credits" or "license" for more information.
>>> import modeller
>>>
If the module 'modeller' loads without giving and error message then you have successfully installed modeller and you should be able to run EasyModeller without any errors.
If you face any problems please use this Google Groups forum to ask questions : Click here
For a detailed tutorial and manual : click here
Tab 1 : Loading Sequence and Template
Tab 2 : Aligning the Templates
Tab 3 : Aligning the Query with Templates
Tab 4 : Generating the model
Additional Functions
1. On the fly MODELLER Code Editing
2. Interactive Ramachandran Plot Viewer
3. Backend code output
4. Automatically displays the model using default PDB viewer
For a detailed tutorial and manual : click here